Overview
In December 2023, GP2 announced the sixth data release on the Terra and the Verily® Workbench platforms in collaboration with AMP® PD. This release includes >20,000 additional participants, adding to the previous releases from the Complex and Monogenic Networks.
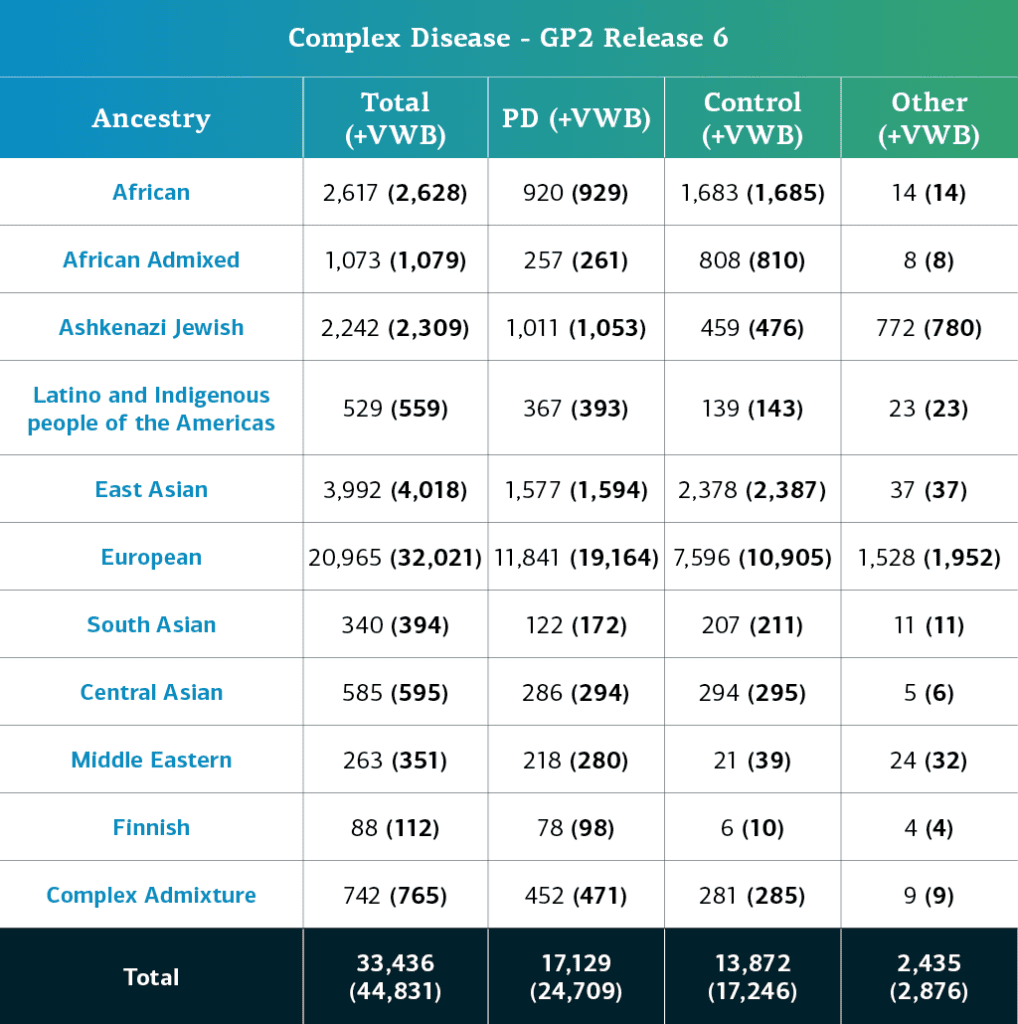
- The complex disease data (genotypes), including locally-restricted samples, now consists of a total of 44,831 genotyped participants (24,709 PD cases, 17,246 Controls, and 2,876 ‘Other’ phenotypes)
- When removing the locally-restricted samples, these now consist of 33,436 (17,129 PD cases, 13,872 Controls, and 2,435 ‘Other’ phenotypes)
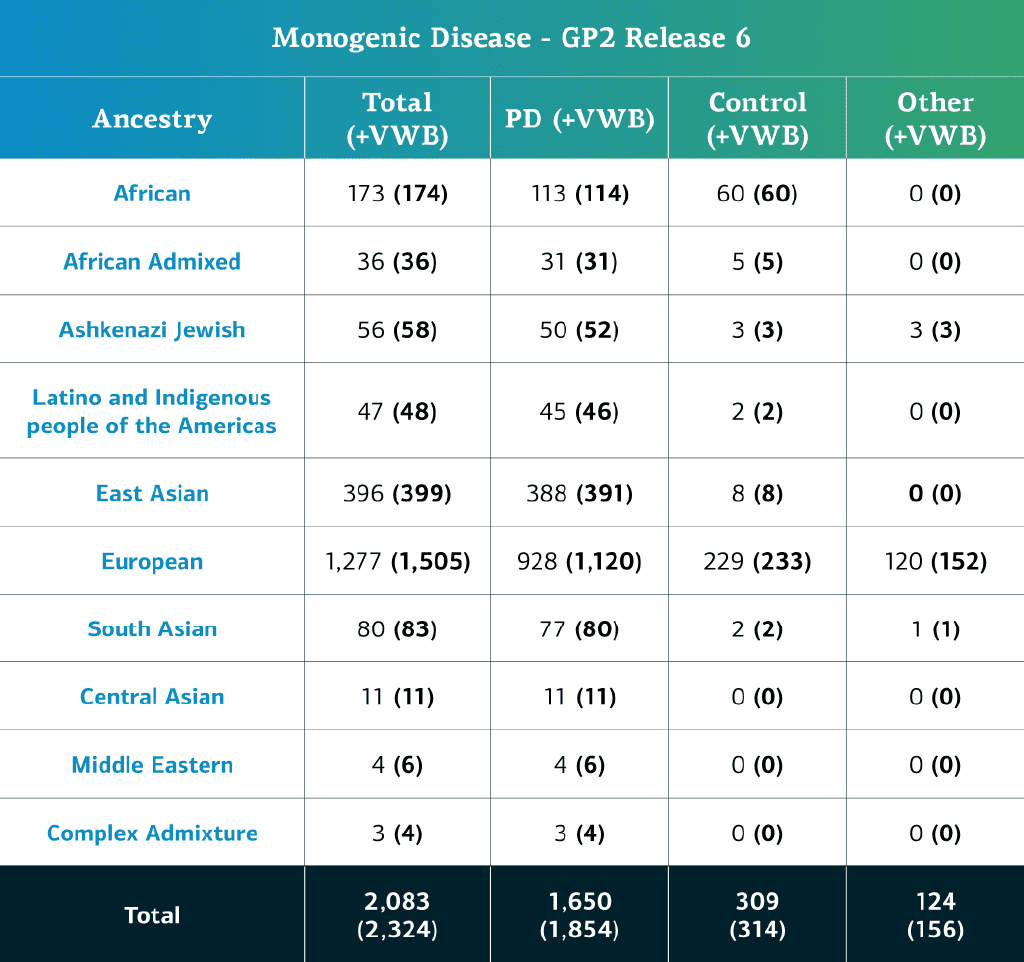
- The monogenic disease data (whole genome sequences) now consists of a total of 2,324 sequenced participants (1,854 PD cases, 314 Controls, and 156 ‘Other’ phenotypes)
- When removing the locally-restricted samples, these now consist of 2,083 (1,650 PD cases, 309 Controls, and 124 ‘Other’ phenotypes)
- 12,585 individuals who have deep clinical phenotyping information also have matching genetic information
What’s New In This Release?
- Additional complex disease (genotyped) and monogenic disease (whole genome) samples
- Introducing locally-restricted GDPR samples via the Verily Viewpoint Workbench
- Introducing clinical data for ~12,000 individuals
- Introducing a new ancestry group → Complex Admixture History (CAH)
- Updates in quality control measures for released genotyping data
- Updates in variant calling, now with DeepVariant, for released whole genome data
Updates in Quality Control
In summary, GenoTools (v1.0.0) performs the following quality control steps: sex mismatches, call rate pruning, checking for duplicates, checking and reporting related individuals, and a heterozygosity rate check.
In contrast to previous releases, we no longer perform the following variant-level filtering: by minor allele frequency (MAF), Hardy-Weinberg (HWE), or minor allele count (MAC). If you would like to filter how we have done for previous releases, we recommend consulting the corresponding README for detailed information and suggested thresholds.
Complex Admixture History (CAH)
CAH, or Complex Admixture History, is a new ancestry group introduced to GP2 for release 6. It was created in response to a large number of samples with South African and other highly admixed individuals being incorrectly predicted as CAS (Central Asian) ancestry in release 5. For release 6, the CAH ancestry group mainly contains samples from Stellenbosch University (Cape Town, South Africa), The Coriell Institute (Camden, New Jersey, United States), and the Parkinson’s Foundation (Miami, Florida, United States). We consider any samples labeled as CAH to be too highly admixed to be included in analyses with other GP2 ancestry groups.
Locality-restricted GDPR samples via the Verily Viewpoint Workbench
We are excited to announce that some users will be able to access locally-restricted samples, otherwise known as samples governed by the General Data Protection Regulation (GDPR) policy, through our collaboration with the Verily Viewpoint Workbench. Workbench is a secure environment for governing and analyzing biomedical data, aimed at enhancing research collaboration and data reproducibility via cloud integration. It supports workspace sharing, including Python and R code, and offers a suite of cloud-native services for data management and analysis. Workbench supports secure data use integration and custom authentication, making it the ideal, secure, and scalable research environment for GP2 to host its locally restricted samples.
At this time, as GP2 continues to roll out data sharing solutions for GDPR-protected data, release 6 will be available to only GP2 consortium members and partners. As testing and implementation continues in early 2024, these solutions will be available to the broader research community. All release 6 samples can be found on Workbench, meanwhile all release 6 samples not governed by GDPR requirements can be found on the community workbench on Terra (like all previous releases). To gain access to the full release on Verily Viewpoint Workbench you must:
- Have approved GP2 Tier 2 access
- Be a GP2 consortium member (contributing cohort, GP2 partner, or project analyses team member)
- Fill out the GDPR-governed sample request form. Upon completing the form, you’ll receive follow up instructions for accessing the Workbench.
Clinical Data
We are also really excited to announce comprehensive deep clinical phenotyping data for 12,585 individuals in this release. This information consists of:
- Age at diagnosis and onset
- Primary, current, and latest diagnoses
- Cognitive exams such as the Mini-Mental State Examination (MMSE) and the Montreal Cognitive Assessment (MoCA)
- Movement Disorder Society-Sponsored Revision of the Unified Parkinson’s Disease Rating Scale (MDS-UPDRS)
- Detailed “other” phenotypes, such as Lewy Body Dementia (LBD)
In this release, each of the 12,585 individuals who have clinical information also have matching genetic information.
Individual-Level Data
We now capture the data from a total of 74 cohorts, 46 are new to this release. Please refer to the GP2 Cohort Dashboard for more information on the cohorts that have been shared.
Genetically-determined ancestry of complex disease GP2 participants is broken into 11 ancestry groups. The table below details the genetically-determined ancestry of complex disease participants in this release that have passed quality control and been imputed. These numbers include samples from previous releases that have been reclustered using the new cluster file and gone through quality control along with the newly genotyped and shared samples unique to this current release.
Variant Calling Whole Genome Sequences using DeepVariant-GLnexus
In this release, in contrast to previous releases, we now use Google’s DeepVariant pipeline coupled with GLnexus for cohort-level variant calling. DeepVariant is a deep learning-based variant caller that outperforms existing state-of-the-art tools by accurately calling individual-level genetic variants. It also simplifies the process, enhancing accuracy and reliability.
Future data releases will continue to grow the diversity of participants available. You can check out our dashboard to see our progress. For users with tier 2 access already, you can explore the data further on our cohort browser, expanded on in a previous blog post.
As always, please refer to the README that accompanies each GP2 release for further details regarding pipelines, data, and analyses!